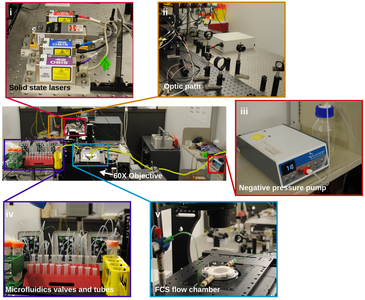
Chromatin structure and transcriptional regulation
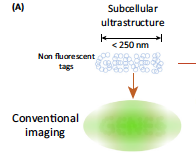
We are interested in the investigation of chromatin structure and at the mechanisms involved in this organization. Our model systems are Drosophila, mice, human, and bacteria. To reach this objective, we develop multiplexed super-resolution and other advanced microscopy methods, as well as gene editing and genome-wide approaches. Below you will find a description of our most recent studies:
Recent papers
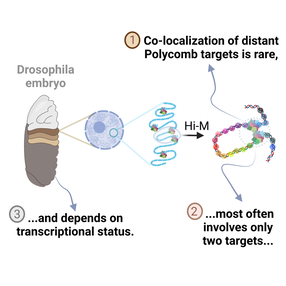
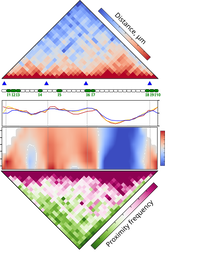
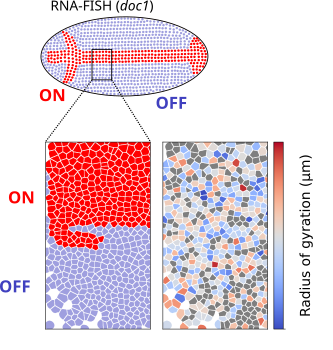
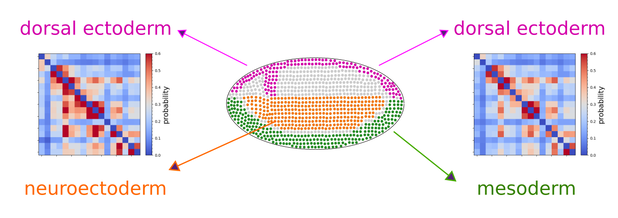
Multiplexed chromatin imaging reveals predominantly pairwise long-range coordination between Drosophila Polycomb genes
|
3D chromatin interactions involving Drosophila insulators are infrequent but preferential and arise before TADs and transcription
|
Multiple parameters shape the 3D chromatin structure of single nuclei
|
Cis-regulatory chromatin loops arise before TADs and gene activation, and are independent of cell fate during development
|
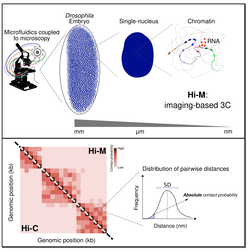
Direct and simultaneous observation of transcription and chromosome architecture in single cells with Hi-M
Andrés M. Cardozo Gizzi, Sergio M. Espinola, Julian Gurgo, Christophe Houbron, Jean-Bernard Fiche, Diego I. Cattoni, Marcelo Nollmann
|
Microscopy-based chromosome conformation capture enables simultaneous visualization of genome organization and transcription in intact organisms
|
TADs Are 3D Structural Units of Higher-Order Chromosome Organization in Drosophila
Quentin Szabo, Daniel Jost, Jia-Ming Chang , Diego I Cattoni, Giorgio L Papadopoulos, Boyan Bonev, Tom Sexton, Julian Gurgo, Caroline Jacquier, Marcelo Nollmann, Frédéric Bantignies, Giacomo Cavalli
Sci Adv 2018 Feb 28;4(2):eaar8082.
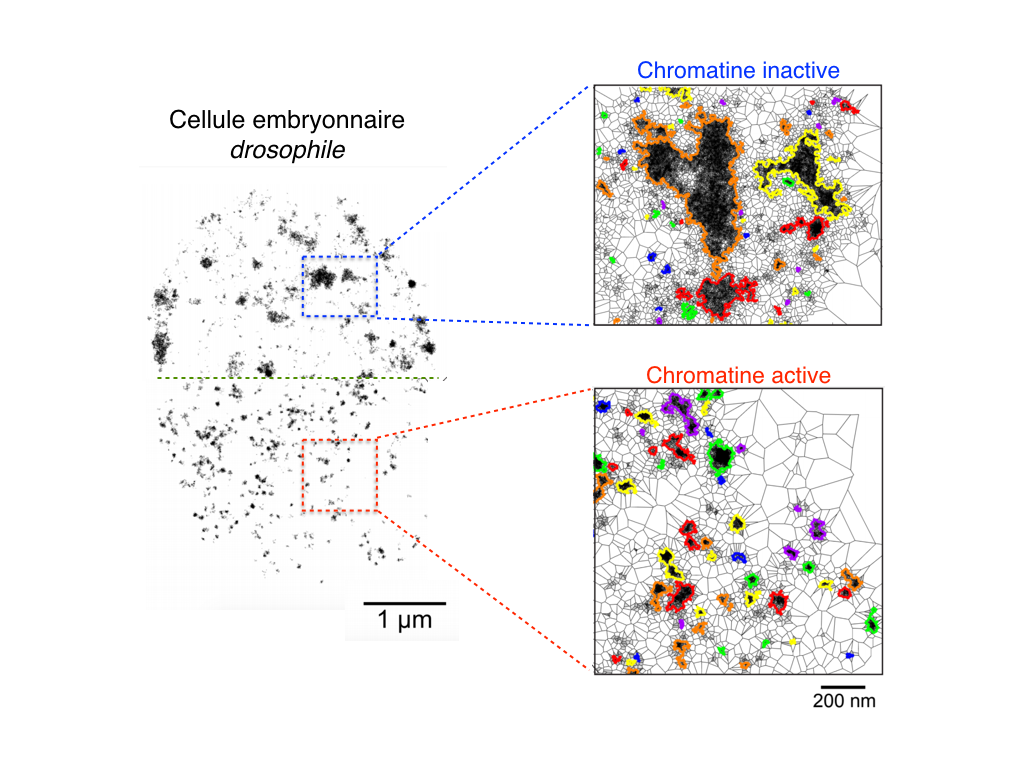
Deciphering the rules of genome folding in the cell nucleus is essential to understand its functions. Recent chromosome conformation capture (Hi-C) studies have revealed that the genome is partitioned into topologically associating domains (TADs), which demarcate functional epigenetic domains defined by combinations of specific chromatin marks. However, whether TADs are true physical units in each cell nucleus or whether they reflect statistical frequencies of measured interactions within cell populations is unclear. Using a combination of Hi-C, three-dimensional (3D) fluorescent in situ hybridization, super-resolution microscopy, and polymer modeling, we provide an integrative view of chromatin folding in Drosophila. We observed that repressed TADs form a succession of discrete nanocompartments, interspersed by less condensed active regions. Single-cell analysis revealed a consistent TAD-based physical compartmentalization of the chromatin fiber, with some degree of heterogeneity in intra-TAD conformations and in cis and trans inter-TAD contact events. These results indicate that TADs are fundamental 3D genome units that engage in dynamic higher-order inter-TAD connections. This domain-based architecture is likely to play a major role in regulatory transactions during DNA-dependent processes.
Single-cell Absolute Contact Probability Detection Reveals Chromosomes Are Organized by Multiple Low-Frequency Yet Specific Interactions
|
Bacterial partition complexes segregate within the volume of the nucleoid
|
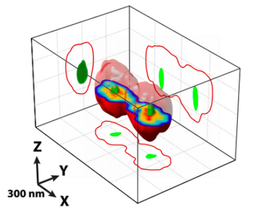
Antoine Le Gall, Diego I. Cattoni, Baptiste Guilhas, Céline Mathieu-Demazière, Laura Oudjedi, Jean-Bernard Fiche, Jérôme Rech, Sara Abrahamsson, Heath Murray, Jean-Yves Bouet, Marcelo Nollmann
Nature Communications 2016; 7:12107 Precise and rapid DNA segregation is required for proper inheritance of genetic material. In most bacteria and archaea, this process is assured by a broadly conserved mitotic-like apparatus in which a NTPase (ParA) displaces the partition complex. Competing observations and models imply starkly different 3D localization patterns of the components of the partition machinery during segregation. Here we use super-resolution microscopies to localize in 3D each component of the segregation apparatus with respect to the bacterial chromosome. We show that Par proteins locate within the nucleoid volume and reveal that proper volumetric localization and segregation of partition complexes requires ATPase and DNA-binding activities of ParA. Finally, we find that the localization patterns of the different components of the partition system highly correlate with dense chromosomal regions. We propose a new mechanism in which the nucleoid provides a scaffold to guide the proper segregation of partition complexes. |
Condensin- and replication-mediated dynamic folding of a bacterial chromosome and its origin domain revealed by super-resolution imaging and Hi-C
Martial Marbouty*, Antoine Le Gall*, Diego I. Cattoni, Axel Cournac, Alan Koh, Jean-Bernard Fiche, Julien Mozziconacci, Heath Murray, Romain Koszul#, Marcelo Nollmann#
Molecular Cell 2015; 59(4):588-602 Chromosomes of a broad range of kingdoms, from bacteria to mammals, are structured by large topological domains, whose precise functional roles and regulatory mechanisms remain elusive. Here, we combine super-resolution microscopies and chromosome-capture technologies to unravel the higher-order organization of the Bacillus subtilis chromosome and its dynamic rearrangements during the cell cycle. We decipher the fine 3D architecture of the origin domain, revealing folding motifs regulated by condensin-like complexes. This organization, along with global folding throughout the genome, is present before replication, disrupted by active DNA replication and re-established thereafter. Single-cell analysis revealed a strict correspondence between sub-cellular localization of origin domains and their condensation state. Our results suggest that the precise 3D folding pattern of the origin domain plays a role in the regulation of replication initiation, chromosome organization and DNA segregation. |
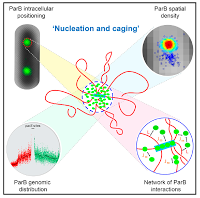
Stochastic Self-Assembly of ParB Proteins Builds the Bacterial DNA Segregation Apparatus
|
Aurore Sanchez,Diego I. Cattoni, Jean-Charles Walter, Jerome Rech, Andrea Parmeggiani, Marcelo Nollmann, and Jean-Yves Bouet
Cell Systems 2015; 1(2):163-73 Many canonical processes in molecular biology rely on the dynamic assembly of higher-order nucleoprotein complexes. In bacteria, the assembly mechanism of ParABS , the nucleoprotein super-complex that actively segregates the bacterial chromosome and many plasmids, remains elusive. We combined super-resolution microscopy, quantitative genomewide surveys, biochemistry, and mathematical modeling to investigate the assembly of ParB at the centromere-like sequences parS . We found that nearly all ParB molecules are actively confined around parS by a network of synergistic protein-protein and protein-DNA interactions. Interrogation of the empirically determined, high-resolution ParB genomic distribution with modeling suggests that instead of binding only to specific sequences and subsequently spreading, ParB binds stochastically around parS over long distances. We propose a new model for the formation of the ParABS partition complex based on nucleation and caging: ParB forms a dynamic lattice with the DNA around parS . This assembly model and approach to characterizing large-scale, dynamic interactions between macromolecules may be generalizable to many unrelated machineries that self-assemble in superstructures |
A matter of scale: how emerging technologies are redefining our view of chromosome architecture.
|
Cattoni DI, Valeri A, Le Gall A, Nollmann M.
Trends Genet. 2015 Aug;31(8):454-64. The 3D folding of the genome and its relation to fundamental processes such as gene regulation, replication, and segregation remains one of the most puzzling and exciting questions in genetics. In this review, we describe how the use of new technologies is starting to revolutionize the field of chromosome organization, and to shed light on the mechanisms of transcription, replication, and repair. In particular, we concentrate on recent studies using genome-wide methods, single-molecule technologies, and super-resolution microscopy (SRM). We summarize some of the main concerns when employing these techniques, and discuss potential new and exciting perspectives that illuminate the connection between 3D genomic organization and gene regulation. |
|
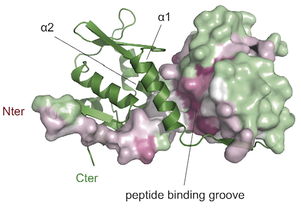
Chromatin insulator factors involved in long-range DNA interactions and their role in the folding of the Drosophila genome
Vogelmann J, Le Gall A, Dejardin S, Allemand F, Gamot A, Labesse G, Cuvier O, Nègre N, Cohen-Gonsaud M, Margeat E, Nöllmann M. PLoS Genetics 2014; 10(8):e1004544 Chromatin insulators are genetic elements implicated in the organization of chromatin and the regulation of transcription. In Drosophila, different insulator types were characterized by their locus-specific composition of insulator proteins and co-factors. Insulators mediate specific long-range DNA contacts required for the three dimensional organization of the interphase nucleus and for transcription regulation, but the mechanisms underlying the formation of these contacts is currently unknown. Here, we investigate the molecular associations between different components of insulator complexes (BEAF32, CP190 and Chromator) by biochemical and biophysical means, and develop a novel single-molecule assay to determine what factors are necessary and essential for the formation of long-range DNA interactions. We show that BEAF32 is able to bind DNA specifically and with high affinity, but not to bridge long-range interactions (LRI). In contrast, we show that CP190 and Chromator are able to mediate LRI between specifically-bound BEAF32 nucleoprotein complexes in vitro. This ability of CP190 and Chromator to establish LRI requires specific contacts between BEAF32 and their C-terminal domains, and dimerization through their N-terminal domains. In particular, the BTB/POZ domains of CP190 form a strict homodimer, and its C-terminal domain interacts with several insulator binding proteins. We propose a general model for insulator function in which BEAF32/dCTCF/Su(HW) provide DNA specificity (first layer proteins) whereas CP190/Chromator are responsible for the physical interactions required for long-range contacts (second layer). This network of organized, multi-layer interactions could explain the different activities of insulators as chromatin barriers, enhancer blockers, and transcriptional regulators, and suggest a general mechanism for how insulators may shape the organization of higher-order chromatin during cell division. |